So I never fully appreciated the value of knowing and assessing the quality of my data until the advent of Next Generation Sequencing. With Sanger sequencing it’s easy enough to load the chromatograms and see the ‘quality’ via peaks yourself and correct anything that is out of the ordinary.
And as we’ve been hearing a lot will determine how well your analysis turns out, sequencing experimental design and of course quality. I have encounter datasets with atrocious quality or missing data that weren’t curated or even looked at in the slightest and then it becomes a monumental effort to go back pull all the raw data and discern what happened…and fix or toss the data out completely (which NO boss wants to hear, but sometimes we have to say).
Junk in = Junk out
Plain and Simple.
Do you trust your sequencer? You shouldn’t, they aren’t perfect as Konrad and others have stated in several lectures and tutorials, data that comes out of the sequencers is not perfect by a long shot and there are a series of filters and quality control measure you really need to take the time to do to assure you are working with good data.
Does this mean you need to turn into a militant about it? No…as Daniel McDonald has stated on many occasions it depends on your ultimate question as to how ‘militant’ you need to be about your QC/QA process. If you are designing site-directed mutagenesis experiements like we do for research ultimately leading to vaccine development and you have to absolutely know every single mutation in a coding sequence and whether it’s real or not and in what abundance (if it’s a polymorphic site) –then yes your inner commandante needs to come out and you need to be sure of the base calling, the read quality and the mapping qualities. However if you are looking at beta diversity, as Daniel has mentioned before, then while you don’t want garbage sequence, your flexibility is increased and perhaps you don’t need to hyperventilate quite so much regarding the quality of every single read and base call in your data.
The same can be inferred for other processes like alignment. As Konrad mentioned…which aligner you use and what parameters you set will depend, how stringent or precise do you have to be with your data? You may have to try a few different aligners and compare. You want an analysis that runs in a reasonable amount of time and has reasonable power to it and uses the majority of the data possible. It is rarely possible to find a program that does it all…perhaps after your python tutorial you will be ready to write one, yes? However, it also depends on the downstream application of the assembly or alignment you create. For instance I work on reference mapping (aligning to reference genome) and curating full genomes of dengue virus and influenza. This is ‘easier’ persay because I have a lovely reference, in fact decades worth of references to draw from to help me but I still get variance in my assemblies when I map or do them de novo, some are better some are worse I generally don’t worry about it though…why? As long as I have complete coverage of the genome and adequate depth and the bases/reads pass my quality filter, all I am interested in is consensus sequence. So as long as I cover the genome per the above, that’s where it ends, as long as I get a reasonable consensus sequence I don’t need to worry so much about every little specific about the aligner/assembler I used. So for my purposes I am militant about sequence and read quality and less concerned about alignment (reference mapping) as long as I don’t have gaps or low coverage regions that will make consensus generation difficult. FYI: I use BWA, generate statistics from a BAM file for every nucleotide position, funnel the BAM to a variant caller, create a VCF (variant call file), eventually read VCF and apply site corrections per the statistics in the VCF > Consensus.fasta
Other downstream programs are also robust against less desirable alignments/assemblies…like UniFrac which you may hear about from Daniel. It is quite robust against crap alignments.
So everything in measure…be aware of the considerations and concerns but do not miss the forest for the trees my fellow bioinformatic minions. Because in the end, if you get so hung up on a particular tree or small subset of random trees then you will miss the beauty of the forest. When you zoom out and look across rolling mountains of trees whose leaves have changed to yellow, gold, red per the condition of autumn –One green tree does not negate the fact that it is autumn…make sense? I hope so.
Alright…I will stop proselytizing and get on with it…
Naiara Rodriguez Ezpeleta
AZTI Tecnalia
Faculty
Topic: Quality Assessment and Control of NGS data
Go through her slides and learn about fasta and fastq and phred scores if you don’t know already. She has a nice exercise that brings back nightmares of math which can be frustrating if you don’t break it down so let me offer some tips to figuring out her Quiz 1 and Quiz 2 slides which are based on the figure below:

Along the bottom of the figure you see a span of characters and letters. They represent a quality score, it’s easier to represent a quality score as a single character as opposed to 2 characters which is what a number score would be. So for instance if you see the ‘!’ character that means the score is 33…
Depending on the platform you will have to subtract from the number in order to obtain the actual quality score. For instance on the figure find Illumina 1.8+ and Sanger, both state +33 so in order to know what the actual score is (0 being bad, 40+ being good) you have to subtract 33. So if a base quality score = ‘!’ it actually has a quality score of 0–so complete crap. For the other platforms you can see on the slide you have to subtract 64. So if you have a base with a score of ‘J’, then by counting it looks like ‘J’ has a score of 74 (notice the I before it has a score of 73, those numbers below the characters are meant to help with counting). BUT you have to subtract 64 IF you’re sequences come from Illumina 1.3, 1.5 or Solexa meaning you actually have a score of 8–once again, complete crap.
You see the long strings of letters at the top, SSSSSSSSS etc, LLLLLLLL etc…those are explained in the figure too, see the letter next to the name of the sequencing platform? Ie. S – Sanger. Well the ‘span’ of those letters is the ‘span’ of characters used to signify the qualities for the that sequencing platform. For example the characters starting at the beginning ‘!’ through the colon character ‘:’ are characters used to describe sequence quality ONLY in Sanger (S) and Illumina (L). Notice that there are a span of characters ‘A’ to ‘I’ where they all overlap meaning if you see qualities that are capital letters between ‘A’ and ‘I’ there’s no way to tell which sequencing platform it came from.
And finally you see the characters at the end…105-126 (lower case ‘i’ to the dash symbol ‘-‘); you’ll notice no Letter designations above those characters…well for the platforms in this figure, they do not use quality scores in this range.
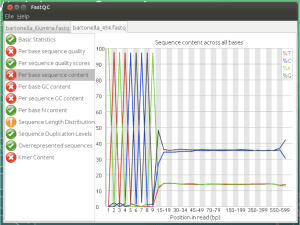
So we used FastQC and FastX-toolkit to look at a variety of datasets and exercises to appreciate how to evaluate our runs as well as pick out common problems that can occur.
Good quality:
Presence of barcodes:
Special cases of left over primers and using special types of DNA like microRNA:
Now in terms of sequence quality you shouldn’t ONLY rely on FastQC, depending on your platform/type of sequencing, sample quality/prep, and overall sequencing results you will have to tweak your protocol. FastQC doesn’t answer everything, it simply gives you and overall snapshot of the quality of your data and points out some ‘concerns’ if there are any.
- What if you have homopolymers or a lot of indels (insertion/deletions)?
- What if you sequenced the wrong organism by accident DOH!
- How do you deal with depth in light of coverage?
- How do you deal with variants?
- Are you even interested in variants or just consensus? Because that’ll affect your overall QC protocol as well…
Lots to think about.
Other cleaning programs mentioned by Naiara and that I’ve heard about include:
- TRIMMOMATIC, a flexible read trimming tool for Illumina sequencing data.
- Galaxy, a web-based genomics pipeline, in which FASTX-Toolkit and FastQC are integrated.
- PRINSEQ, either as a standalone package or through a web-interface can generate summary statistics of sequence and quality data, which can subsequently be used to filter, reformat and trim next-generation sequence data.
- Perl and Bioperl, to write small scripts. There already exist a very large number of packages devoted to genomics in Bioperl.
- R and Bioconductor, are other solution to import and verify data. There are many contributed packages and modules available.
- MUSKET: An efficient multistage k-mer-based corrector for Illumina short-read data
- ReadClean (RC454): If you are dealing with 454 data
- QUAKE: Correct substitution sequencing errors in experiments with deep coverage (e.g. >15X), specifically intended for Illumina sequencing reads
- Hammer: Error-correction of high-throughput sequencing datasets
- SEECER: RNA-seq error correction
Many of these programs use a Kmer-spectrum approach to error correction, a good slide show that illustrates what this is was presented by Srinivas Aluru from Iowa State University.
And for those interested in more context with respect to error correction in transcriptomes, Drs. Eisen and MacMains have a manuscript up on arXiv preprint server entitled: Improving transcriptome assembly through error correction of high-throughput sequence reads.
So in sum…
- Be aware of your sequence quality
- Know what tools are available
- Know how to fix common problems in your data
But in the end…you need to find that balance, assessing your question and the end goal of your study and try not to
Miss the forest for the trees…
…Dr. Mel